 See gallery
See gallery
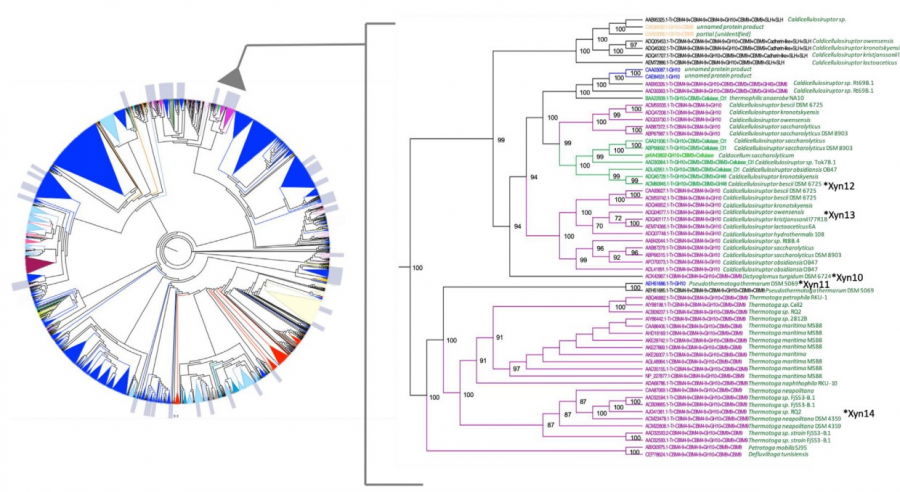
Cladogram of the GH10 family
2021-05-28
New scientific paper on xylanase Xyn11, active at extreme conditions of temperature and alkalinity
One of the initial objectives of the European Project WoodZymes (www.woodzymes.eu) was to obtain a xylanase biocatalyst active at extreme conditions of temperature and alkaline pH, suitable for use in the pulp and paper industry. An enzyme with the desired characteristics, named Xyn11 has been produced. To achieve this objective, a bioinformatic analysis of family 10 of glycoside hydrolases (GH10), one of the major families of protein sequences that contain xylanases, was performed. More than 2000 protein sequences that are annotated in databases were analyzed.
The great majority of these sequences have been obtained as a result of massive and metagenomic sequencing analyses. Based on their homology to known enzymes, these proteins are supposed to have a particular enzymatic activity but less than 1% of them have been biochemically characterized. Bioinformatic analysis allows to pinpoint individual sequences that probably have prefixed properties, narrowing down the number of candidates susceptible of undergoing experimental analysis.
A cladogram (i.e. a diagram showing genetic relatedness) of the over 2000 GH10 family sequences was constructed, in which a group potentially corresponding to thermophilic and alkaliphilic xylanases (labelled TAK cluster) could be mapped. From this cluster, five sequences were chosen for biochemical analysis. Synthetic genes encoding these proteins, with genetic code optimized for expression in E. coli, were obtained. The proteins resulting from expressing the synthetic genes cloned in E. coli were produced, purified and their enzymatic activity was assayed at different conditions of pH and temperature.
Among these, Xyn11, which corresponds to a protein sequence from the thermophilic bacterium Pseudothermotoga thermarum, showed prominent xylanase activity at 90 oC and pH 10.5. Moreover, the addition to Xyn11 of a carbohydrate binding protein domain from the thermophilic bacterium Pyrococcus furiosus, further enhanced the enzymatic activity under extreme conditions.
This work has been carried out at the Laboratory of Enzyme Structure and Function, from the Institute of Agrochemistry and Food Technology, Spanish Research Council (CSIC) lead by Dr. Julio Polaina. In collaboration with the group lead by Dr. Julia Sanz-Aparicio at the Department of Crystallography of the Institute Rocasolano of Physical Chemistry (CSIC), the three-dimensional structure of Xyn11 has been solved to a resolution of 1.8 Å. This allowed to define the atomic interactions responsible for the extreme properties of this enzyme.
 See gallery
See gallery


